pacman::p_load(dplyr,
tidyr,
broom,
stringr,
modelsummary,
tinyplot,
ggplot2)Likelihood comparison*
Goals
The goal of this aside is to introduce you to some concepts that the book doesn’t really address until the very end: mainly likelihood-based model selection and the generalized linear model. Since likelihood came up in at the end of the Section A Recap and we’re now dealing with more realistic models (Chapter 6), this seemed like a good time.
Set up
As usual, we’ll set packages and our theme.
tinytheme("ipsum",
family = "Roboto Condensed",
palette.qualitative = "Tableau 10",
palette.sequential = "agSunset")theme_set(
theme_minimal(base_family = "Roboto Condensed") +
theme(panel.grid.minor = element_blank())
)
tableau10 <- c("#5778a4", "#e49444", "#d1615d", "#85b6b2", "#6a9f58",
"#e7ca60", "#a87c9f", "#f1a2a9", "#967662", "#b8b0ac")
options(
ggplot2.discrete.colour = \() scale_color_manual(values = tableau10),
ggplot2.discrete.fill = \() scale_fill_manual(values = tableau10)
)Get the same data we’ve been using.
state.x77_with_names <- as_tibble(state.x77,
rownames = "state")
states <- bind_cols(state.x77_with_names,
region = state.division) |> # both are in base R
janitor::clean_names() |> # lower case and underscore
mutate(south = as.integer( # make a south dummy
str_detect(as.character(region),
"South")))Evaluating likelihoods
50 cases isn’t very many but it will be even easier to understand what we’re doing with an even smaller sample. So let’s restrict the sample to the southern US states.
s_states <- states |>
filter(south == 1)Null model likelihood
Here’s the prediction of the “null model”:
Data prep
s_states$mean_lex <- mean(s_states$life_exp)
width <- sd(s_states$life_exp / sqrt(nrow(s_states))) * qt(.995, 49) # 99%Show code
plt(mean_lex ~ hs_grad,
data = s_states,
ymax = mean_lex + width,
ymin = mean_lex - width,
ylim = c(67, 73),
type = "ribbon",
main = "Life expectancy and education",
sub = "US states, 1977",
xlab = "% high school graduates",
ylab = "Life expectancy at birth")
plt_add(life_exp ~ hs_grad,
data = s_states,
type = "p")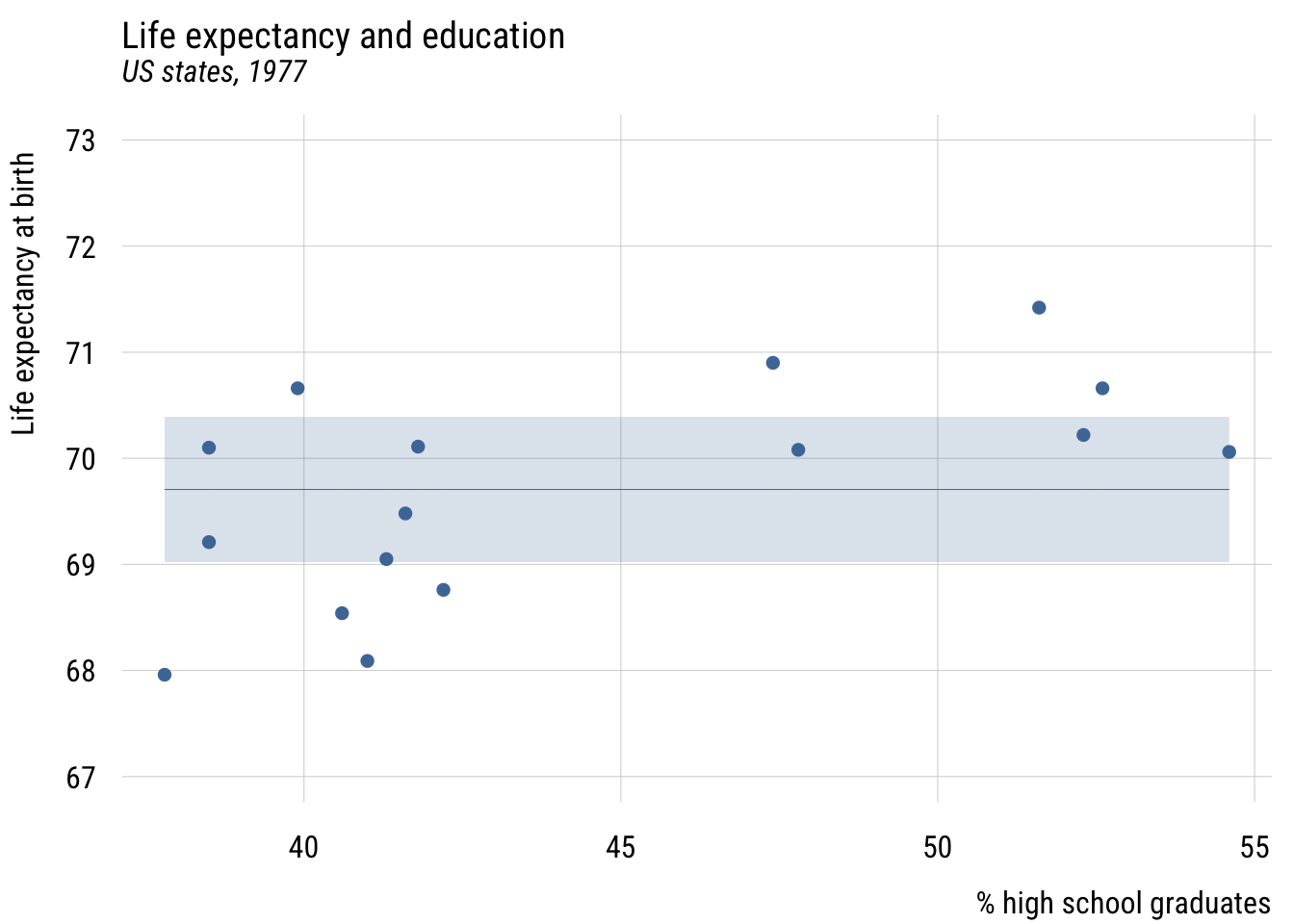
Show code
ggplot(s_states, aes(x = hs_grad)) +
geom_ribbon(aes(ymin = mean_lex - width, ymax = mean_lex + width),
fill = tableau10[1], alpha = 0.2) +
geom_segment(x = min(s_states$hs_grad), xend = max(s_states$hs_grad),
y = unique(s_states$mean_lex), yend = unique(s_states$mean_lex),
color = tableau10[1]) +
geom_point(aes(y = life_exp), color = tableau10[1]) +
coord_cartesian(ylim = c(67, 73)) +
labs(
title = "Life expectancy and education",
subtitle = "US states, 1977",
x = "% high school graduates",
y = "Life expectancy at birth")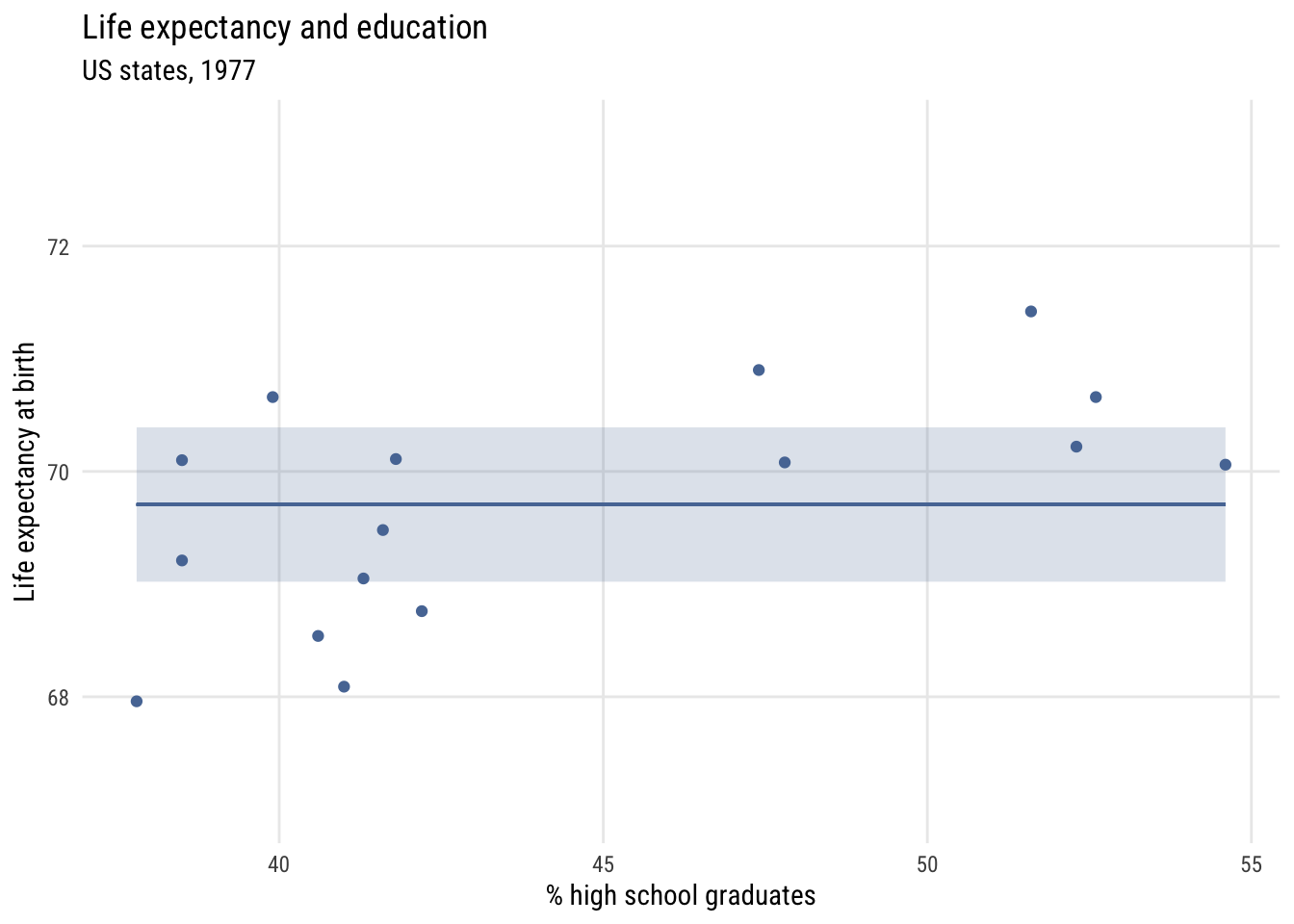
We can write this null model like this, which is a little different than we see in the book. But it will generalize well to other types of models.
\[\begin{align} \text{lifeexp}_i &\sim \mathcal{N}(\mu_i, \sigma) \\ \mu_i &= \beta_0 \end{align}\]
This representation allows for a direct representation of the likelihood, which we saw in the Section A Recap. Sometimes this representation is called the generalized linear model, which is composed of two parts:
- A distributional family for the outcome
- A linear predictor that links conditional expectations to the key parameter in that family (here, \(\mu_i\), or the conditional mean)
We can calculate get the likelihood of the model given the data either the easy way…
mc <- lm(life_exp ~ 1, data = s_states)
logLik(mc)'log Lik.' -22.53801 (df=2)Or we can get it the hard way…
m_hat <- mc$coefficients[1] # prediction of outcome for all cases
s_hat <- sd(mc$residuals) # estimate of sigma
s_states <- s_states |>
mutate(l_mc = dnorm(life_exp, m_hat, s_hat), # row-wise likelihood
ll_mc = log(l_mc)) # row-wise log-likelihoodHere are the likelihoods:
[1] 0.31759577 0.25254287 0.36759389 0.25254287 0.20356969 0.36237192
[7] 0.25427082 0.34397205 0.11181475 0.34689364 0.09572509 0.09071031
[13] 0.36099167 0.19734482 0.36504343 0.38083463Here are the log-likelihoods (which we actually use):
[1] -1.1469759 -1.3761743 -1.0007765 -1.3761743 -1.5917469 -1.0150842
[7] -1.3693554 -1.0671949 -2.1909118 -1.0587371 -2.3462748 -2.4000842
[13] -1.0189004 -1.6228027 -1.0077389 -0.9653901And here’s the sum, which is the model log-likelihood.
sum(s_states$ll_mc)[1] -22.55432Augmented model likelihood
As before, we will add the predictor here so that we make different predictions for different cases.
Show code
plt(life_exp ~ hs_grad,
data = s_states,
ylim = c(67, 73),
main = "Life expectancy and education",
sub = "US states, 1977",
xlab = "% high school graduates",
ylab = "Life expectancy at birth")
plt_add(type = "lm",
level = .99)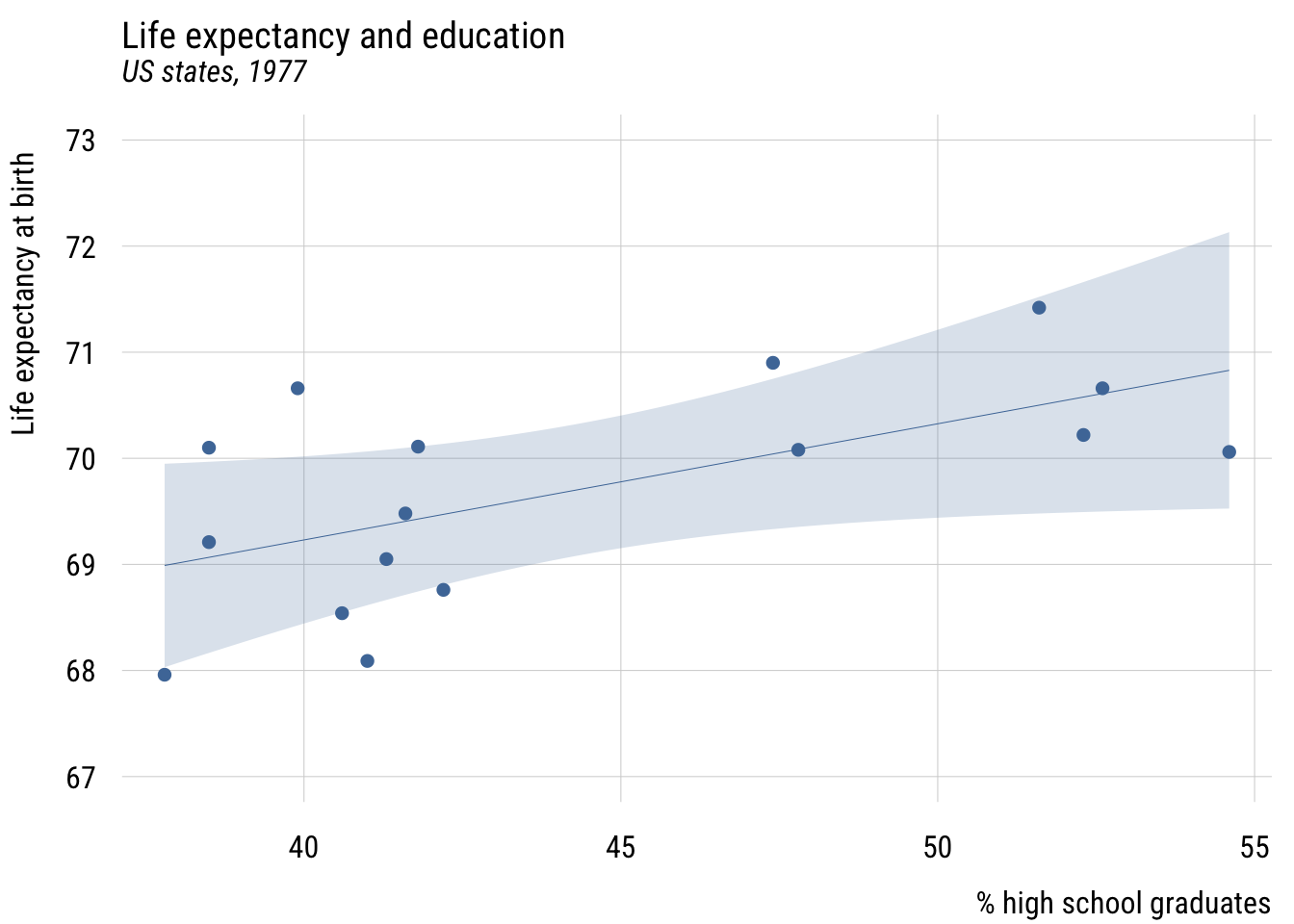
Show code
ggplot(s_states, aes(x = hs_grad, y = life_exp)) +
geom_point(color = tableau10[1]) +
geom_smooth(method = "lm", formula = y ~ x, level = .99,
color = tableau10[1], fill = tableau10[1], alpha = 0.2) +
coord_cartesian(ylim = c(67, 73)) +
labs(
title = "Life expectancy and education",
subtitle = "US states, 1977",
x = "% high school graduates",
y = "Life expectancy at birth")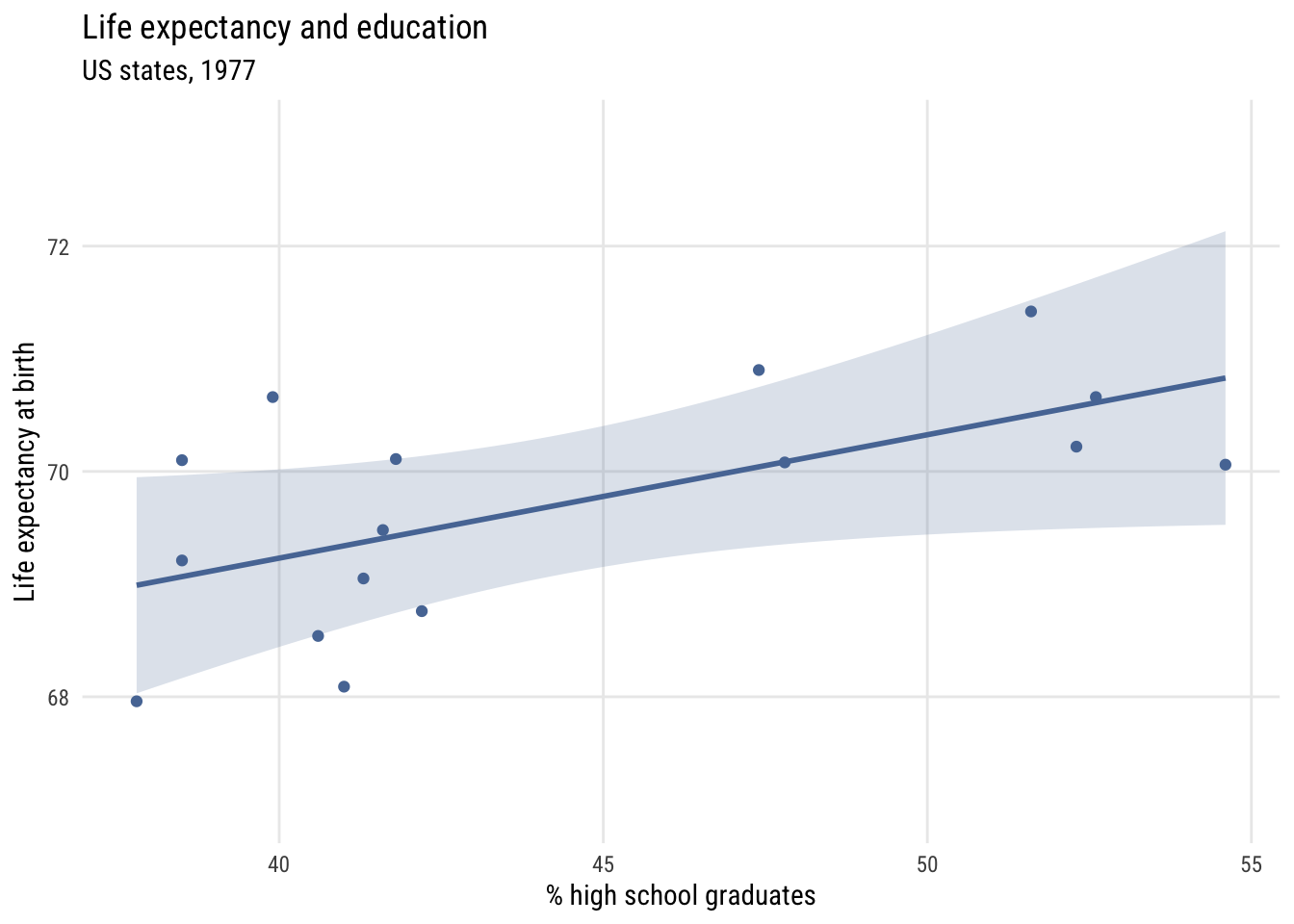
We can write this model like this:
\[\begin{align} \text{lifeexp}_i &\sim \mathcal{N}(\mu_i, \sigma) \\ \mu_i &= \beta_0 + \beta_1\,\text{hsgpct}_i \end{align}\]
And estimate it like this:
ma <- lm(life_exp ~ hs_grad, data = s_states)And again we can get the row-wise likelihoods.
Show code
m_hat <- predict(ma)
s_hat <- sd(ma$residuals)
s_states <- s_states |>
mutate(l_ma = dnorm(life_exp, m_hat, s_hat), # row-wise likelihood
ll_ma = log(l_ma)) |> # row-wise log-likelihood
ungroup()
s_states$l_ma [1] 0.4567596 0.1002810 0.3139155 0.4940019 0.3186773 0.2175253 0.3352283
[8] 0.4486153 0.1486593 0.4871563 0.2582879 0.2188331 0.3459233 0.2804524
[15] 0.4949372 0.4928553And even compare them…
Show code
s_states$l_ma / s_states$l_mc [1] 1.4381791 0.3970850 0.8539736 1.9561111 1.5654455 0.6002820 1.3183908
[8] 1.3042202 1.3295143 1.4043392 2.6982255 2.4124395 0.9582585 1.4211289
[15] 1.3558310 1.2941451This is how much more (or less) likely each point is given Model A’s predictions compared to Model C’s. You can see that most get better although a few do get worse.
Finally, let’s look at the log-likelihood.
logLik(ma)'log Lik.' -18.73661 (df=3)You can see that the log-likelihood of Model A (-18.74) is higher/better than the log-likelihood of Model C (-22.54). But how can we interpret this comparison?
ll_diff <- as.numeric(logLik(ma)) - as.numeric(logLik(mc))
ll_diff # the difference[1] 3.8014exp(ll_diff) # the exponentiated difference[1] 44.76381Exponentiating the log-likelihood difference tells us that Model A is about 45 times more likely than Model C given the data. That seems like a lot! But remember that any more complicated model is going to be at least somewhat more likely. So what is a more systematic way to compare them?
Comparing model likelihoods
We are going to consider two ways to compare likelihoods in this section: the likelihood ratio test and the proportional reduction in deviance. Then we’ll talk about some other ways in the next section.
Likelihood ratio test
It turns out that two times the log-likelihood difference can be used as a test statistic (often called \(G^2\)) for testing the null hypothesis that any parameters you added could be set to zero1.
1 In this case, of course, we only added one parameter, so there are several ways we could test this null hypothesis. These include the \(t\)- or (1 numerator df) \(F\)-test that you’ve seen already. But there are others.
2 We might point out that \(t\) is to \(F\) as \(z\) is to \(\chi^2\). We could also say that \(t\) is to \(z\) as \(F\) is to \(\chi^2\). Both \(t\) and \(F\) are “small sample” statistics whereas \(z\) and \(\chi^2\) are “large sample” or “asymptotic” statistics. The advantage of these asymptotic statistics is that they generalize to almost every type of statistical model, not just linear regression.
This is almost exactly like the \(F\) test that you know already. The only difference is looking up an \(F\) statistic on an \(F\) table, you look up the \(G^2\) statistic on the \(\chi^2\) (“chi-square”) table with \(df\) equal to the number of parameters that differ between the models (just like numerator \(df\) in the \(F\) distribution)2.
\[ G^2 = 2 \bigl[ \ell(\hat{\theta_1}) - \ell(\hat{\theta_0}) \bigr] \overset{\text{approx}}{\sim} \chi^2_{df} \]
Data prep
x <- seq(0, 10, length.out = 300)
y <- dchisq(x, df = 1)Show code
plt(y ~ x,
type = "l",
sub = expression(chi[(1)]^2 ~ ": critical (99%, dashed) and observed values"),
xlab = expression(G^2 ~ "test statistic"),
ylab = "Density",
xlim = c(0, 10),
lwd = 2)
abline(v = qchisq(.99, 1), lty = 2, lwd = 2)
abline(v = 2*ll_diff, col = "#E69F00", lwd = 2)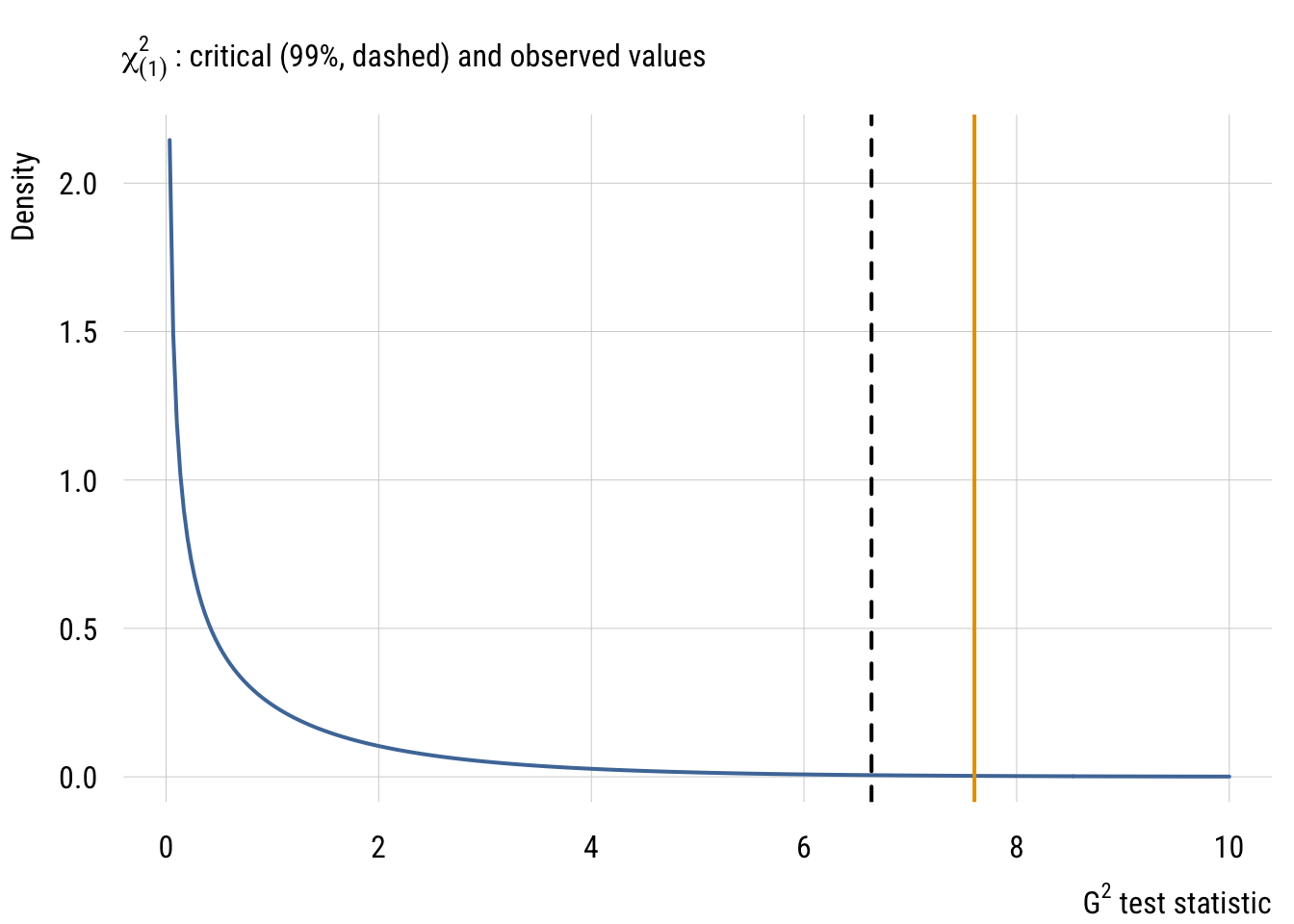
Show code
ggplot(data.frame(x = x, y = y), aes(x = x, y = y)) +
geom_line(linewidth = 1, color = tableau10[1]) +
geom_vline(xintercept = qchisq(.99, 1), linetype = "dashed", linewidth = 1) +
geom_vline(xintercept = 2 * ll_diff, color = "#E69F00", linewidth = 1) +
coord_cartesian(xlim = c(0, 10)) +
labs(
subtitle = expression(chi[(1)]^2 ~ ": critical (99%, dashed) and observed values"),
x = expression(G^2 ~ "test statistic"),
y = "Density")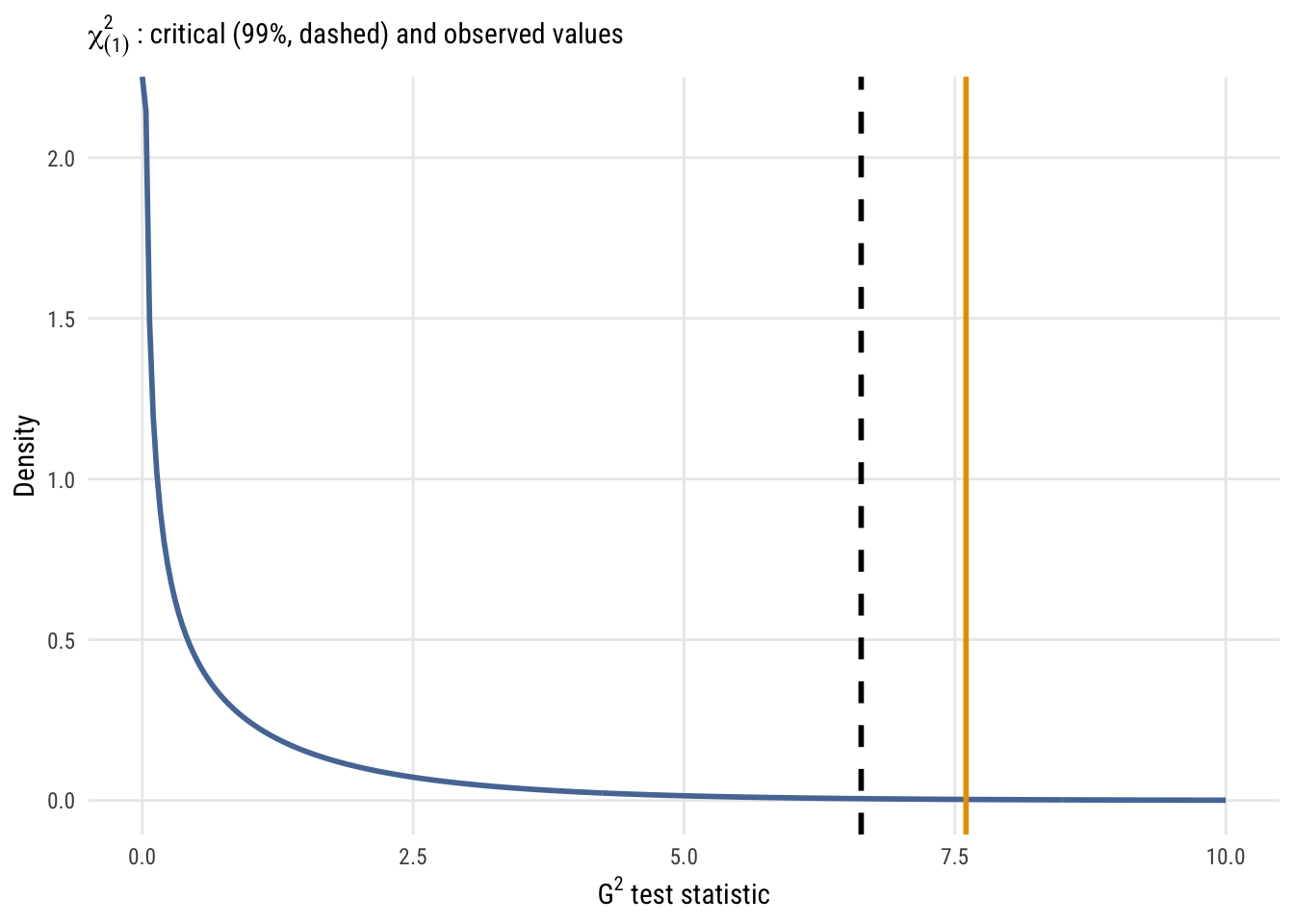
The resulting p-value here is 0.00583, which is smaller than what we’d get from the same \(F\) test (0.01123), although both values are pretty small. The former is smaller is because the \(\chi^2\) is essentially an \(F\) distribution with infinite denominator degrees of freedom, so there is less mass in the tail of the distribution. In other words, weird stuff (i.e., “right tail” stuff) is less likely to occur as a process goes to infinity.
To easily get the likelihood-ratio test (LRT) for these models in R, you might want to use the lmtest::lrtest() function3.
3 With pretty much everything except lm-class objects, the anova() function (e.g, anova(mc, ma)) will give you the LRT. For lm objects, however, you get the \(F\) test by default. And if you add the argument test = "LRT" or test = "Chisq" to the anova() call, you will, very confusingly, not be given the LRT but the Wald \(\chi^2\) test (which is not the same thing as the LRT but rather looking up the \(F\)-statistic on the \(\chi^2\) reference distribution).
lmtest::lrtest(mc, ma)Likelihood ratio test
Model 1: life_exp ~ 1
Model 2: life_exp ~ hs_grad
#Df LogLik Df Chisq Pr(>Chisq)
1 2 -22.538
2 3 -18.737 1 7.6028 0.005828 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1This is the same as what we got doing it by hand above.
Proportional reduction of deviance
Deviance (\(D\)) is a term that generally means -2 times the log-likelihood (\(\ell\)) of a model. In other words:
\[ D_{\text{M}} = -2\ell_{\text{M}} \]
The deviance is a likelihood-based a way of answering the question “how far away are the predictions of this model from the data?” Lower deviance is better. This is in spirit the likelihood version of the sum-of-squares, where lower is also better.
This can be confusing because the function deviance() used on an lm object will return the residual sum of squares (RSS or SSE) which is NOT the same thing as \(-2\ell\).4 To prove it, let’s compare the likelihood-based deviance of model A with its RSS.
4 Even though it’s confusing, it at least makes some sense because the RSS is a measure of how far away the model predictions are from the data. Which is, of course, what deviance means in spirit.
dev_a <- -2 * as.numeric(logLik(ma))
dev_a # this is D = -2LL[1] 37.47323rss_a <- sum(ma$residuals^2)
rss_a # this is RSS [same as deviance(ma) here, confusingly!][1] 9.745332You can see they are not the same!
In Chapter 15 of the book, the authors talk about PRD as:
\[ PRD = \frac{D_C - D_A}{D_C} \]
This is reasonable for some other models, but it won’t work properly for linear models for technical reasons5, so it’s better to use a different formula:
5 It’s this pesky \(\sigma\) estimate that is causing the problems. Deviance means comparing models to a theoretical “perfect” (or “saturated”) model. With linear regression, that’s an RSS of zero. But that doesn’t make sense with likelihood-based deviance here because as RSS goes to zero, \(\sigma\) goes to zero, which means likelihood goes to infinity. So we can’t directly use the ratio of likelihoods without taking the difference first (which cancels the “saturated” likelihoods out of the equation).
First we need the “deviance drop” or difference in deviance between the null and augmented model:
\[ \Delta D = D_C - D_A \]
Then we can use that \(\Delta D\) in the equation below (as long as Model C is a null, i.e., intercept-only, model).
\[ R^2 = \frac{\text{RSS}_C - \text{RSS}_A}{\text{RSS}_C} = 1 - \exp \left( \frac{-\Delta D}{n} \right) \]
As implied by this equality, either formula will give exactly the same \(R^2\) for a linear regression model6.
6 The \(\Delta D\) version is called the Cox-Snell pseudo \(R^2\), although it’s not really “pseudo” for linear regression since it’s exactly the same as the good-old-fashioned \(R^2\). There are other pseudo \(R^2\) values you might learn about someday, but they aren’t important right now.
dev_c <- -2 * as.numeric(logLik(mc))
rss_c <- sum(mc$residuals^2)
1 - exp( -(dev_c - dev_a ) / nrow(s_states)) # R^2: using diff in -2LL[1] 0.3782238(rss_c - rss_a) / rss_c # R^2: using RSS[1] 0.3782238You can confirm that these are the same.